Cells - How Automated Counting and Sizing Data make Mammosphere Assays more Practical and Reliable
Introduction
To fully understand the mechanisms underlying breast cancer, research often relies heavily on functional assays provided by cellular models. Two cellular models that have become more pervasive for breast cancer researchers in the last 10 years are mammosphere and breast tumorsphere culture. Mammospheres are cultured from primary breast tissue or cell lines, while tumorspheres are cultured from primary tumor cells and cancer cell lines from multiple tissues. Mammosphere and tumoursphere assays are widely used in vitro to identify prospective cancer-initiating stem cells that can propagate clonally to form spheres in free-floating conditions.
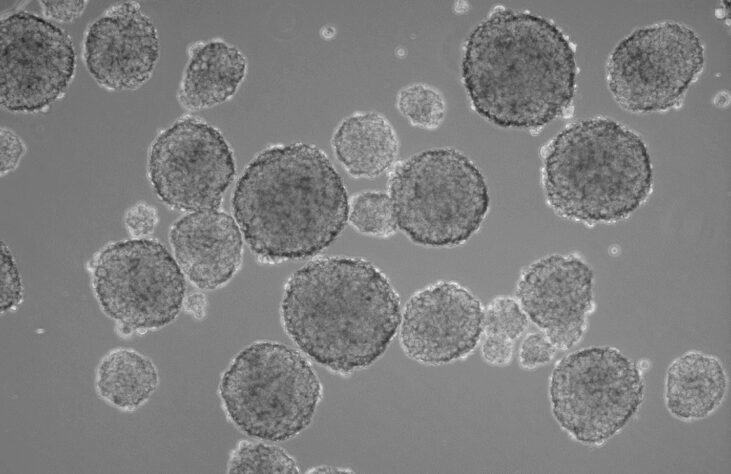
A classic view of mammospheres under low magnification, phase contrast microscopy
Researchers do tend to be interested 3D sphere cultures over its adherent counterpart, as growing cells in 3D provides higher physiological relevance than traditional monolayer cultures. The mammosphere culture system involves seeding mammary epithelial cells at low density in an environment that prevents their adherence to a substratum and enables their proliferation in suspension as spherical clusters.
A multitude of the applications today using mammospheres look directly at kinetic cellular growth of these spheres in different situations. These applications include the toxicity of specific drugs, microenvironment manipulation, and even comparing different cancer phenotypes to one another. These studies always come with a multitude of challenges with one of the most obvious being how to efficiently and precisely count and size 10’s-100’s of wells containing small spheres across an entire study.
The most common way researchers seem to count these spheres today is manually, using bright field microscopy. While this certainly works there are several issues researchers do run into. Please note this topic on microscope counting is covered in greater depth in this article but the most salient points are:
- Microscopes can only look at one plane at a time and counting in 3D means a skilled researcher is looking across not only the X and Y axes but also the Z axis to identify and count spheres.
- It requires a great deal of training and practice to be able to obtain correct counts and become efficient.
- The task itself is time consuming, using up valuable time for what is essentially counting circles.
- Whether wittingly or unwittingly, bias of counts is always a concern.
- Research has shown that both intra- and inter-observer variability is present in counts even among the highly trained.
- Last, unless using a confocal microscope which is time consuming and expensive, there is no way through manual counting to garner any meaningful sizing information on spheres. Sizing and other statistical parameters can be helpful when looking at toxicology, for example, as the growth rate of tumor spheres is also predictive of a treatment’s significance.
The way forward: The GelCount detects and measures mammospheres and tumorspheres in 3D culture

The GelCount: intended for processing adherent or 3D mammosphere colony formation assays
The GelCount is an integrated hardware and PC software platform developed by Oxford Optronix for automated high throughput imaging and analysis of mammalian cell colonies (both adherent and non-adherent). Whether it be the counting and sizing of spheroids, tumor forming, organoid, or clonogenic assays, the GelCount has proven to be a highly capable system for this task. As such, the system is found in labs world-wide and has been cited in over 300 publications (PDF here).
The GelCount differs from other counting approaches. After it takes its image, researchers can use the software to define specific thresholds (including minimum and maximum sizes) to be able to count exactly what’s needed. These thresholds can then be automated to run across an entire study, counting each well or Petri dish with the exact same user defined specifications, meaning there will be no variability in counts. Additionally, the systems learning curve is minimal and anyone in the lab can be taught how to use the system in a short period of time. From a throughput standpoint, on average it takes the GelCount about 12 minutes to count and analyze four (4) separate 6-well plates of 3D mammospheres imaged at high resolution.
Since the GelCount can count objects in 3D space (up to 5 mm depth) it has all the capability needed to image and resolve most mammosphere applications. All the issues seen in the microscope counting techniques stated above (time consuming, counter reliability, low throughput, observer bias), the GelCount entirely removes and will almost certainly provide a more consistent count while increasing throughput across an entire experiment. Crucially the system also intrinsically measures mammospheres diameter and calculated volume which, as explained earlier, can make a simple counting experiment far more dynamic.
Last and likely of interest to many researchers, while many labs do have their own unit, the GelCount is an ideal shared facility. It is often accommodated in a central location within a department or in a core facility. Furthermore, rather than occupy the device, users can take the images generated and transfer them to a separate workstation (with the software installed) for the analysis stage, freeing the imager for the next user.
Mammosphere publications using the GelCount with brief description
As the GelCount has been a staple in cancer research labs and institutions there are a plethora of papers citing mammospheres that have successfully utilized the GelCount for their assays. Below are a few select examples.
Fitzpatrick et al. 2017
Robotic Mammosphere Assay for High-Throughput Screening in Triple-Negative Breast Cancer. Society for Laboratory Automation and Screening
Used the GelCount and demonstrates how this system can contribute to high throughput assays
Hu et al. 2021
Decorin-mediated suppression of tumorigenesis, invasion, and metastasis in inflammatory breast cancer. Communications Biology
Used the GelCount to count colonies (Fig 1B, 1G, 4B) and first and second generation Mammospheres (Fig 1E, 1F, 4E, 4F)
Gagliardi et al. 2020
Differential functions of ERK1 and ERK2 in lung metastasis processes in triple-negative breast cancer. Scientific Reports
Used the GelCount to count colonies (Fig 2D) and Mammospheres (Fig 3A)
Castro et al. 2019
Sulforaphane Suppresses the Growth of Triple-negative Breast Cancer Stem-like Cells In vitro and In vivo. Cancer Prevention Research
Used the GelCount to count and size mammospheres (Fig 2A-D, 3B-F)
Bhat et al. 2018
GPCRs profiling and identification of GPR110 as a potential new target in HER2+ breast cancer. Breast Cancer Research and Treatment
Used the GelCount to count colonies (Fig 7) and first and second generation Mammospheres (Fig 8)
--------------------------------------------





